
Nextflow and nf-core mentorship rocket.
With the third round of the Nextflow and nf-core mentorship program now behind us, it’s time to pop the confetti and celebrate the outstanding achievements of our latest group of mentors and mentees!
As with the first and second rounds of the program, we received hundreds of applications from all over the world. Mentors and mentees were matched based on compatible interests and time zones and set off to work on a project of their choosing. Pairs met regularly to work on their projects and reported back to the group to discuss their progress every month.
The mentor-mentee duos chose to tackle many interesting projects during the program. From learning how to develop pipelines with Nextflow and nf-core, setting up Nextflow on their institutional clusters, and translating Nextflow training materials into other languages, this cohort of mentors and mentees did it all. Regardless of all initial challenges, every pair emerged from the program brimming with confidence and a knack for building scalable and reproducible scientific workflows with Nextlfow. Way to go, team!
Participants of the third round of the mentorship program.
Abhay Rastogi is a Clinical Research Fellow at the All India Institute Of Medical Sciences (AllMS Delhi). During the program, he wanted to contribute to the nf-core/sarek pipeline. He was mentored by Matthias De Smet, a Bioinformatician at the Center for Medical Genetics in the Ghent University Hospital. Together they worked on developing an nf-core module for Exomiser, a variant prioritization tool for short-read WGS data that they hope to incorporate into nf-core/sarek. Keep an eye out for this brand new feature as they continue to work towards implementing this new feature into the nf-core/sarek pipeline!
Alan Möbbs, a Bioinformatics Analyst at MultiplAI, was mentored by Simon Pearce, Principal Bioinformatician at the Cancer Research UK Cancer Biomarker Centre. During the program, Alan wanted to create a custom pipeline that merges functionalities from the nf-core/rnaseq and nf-core/rnavar pipelines. They started their project by forking the nf-core/rnaseq pipeline and adding a subworkflow with variant calling functionalities. As the project moved on, they were able to remove tools from the pipeline that were no longer required. Finally, they created some custom definitions for processing samples and work queues to optimize the workflow on AWS. Alan plans to keep working on this project in the future.
Cen Liau is a scientist at the Bragato Research Institute in New Zealand, analyzing the epigenetics of grapevines in response to environmental stress. Her mentor was Chris Hakkaart, a Developer Advocate at Seqera. They started the program by deploying the nf-core/methylseq pipeline on New Zealand’s national infrastructure to analyze data Cen had produced. Afterward, they started to develop a proof of concept methylation pipeline to analyze additional data Cen has produced. Along the way, they learned about nf-core best practices and how to use GitHub to build pipelines collaboratively.
Chenyu Jin is a Ph.D. student at the Center for Palaeogenetics of the Swedish Museum of Natural History. She worked with Ben Sherman, a Software Engineer at Seqera. Together they worked towards establishing a workflow for recursive step-down classification using experimental Nextflow features. During the program, they made huge progress in developing a cutting-edge pipeline that can be used for analyzing ancient environmental DNA and reconstructing flora and fauna. Watch this space for future developments!
Georgie Samaha, a bioinformatician from the University of Sydney, was mentored by Cristina Tuñi i Domínguez, a Bioinformatics Scientist at Flomics Biotech SL. During the program, they developed Nextflow configuration files. As a part of this, they built institutional configuration files for multiple national research HPC and cloud infrastructures in Australia. Towards the end of the mentorship, they built a tool for building configuration files that they hope to share widely in the future.
Ícaro Maia Santos is a Ph.D. Candidate at the University of São Paulo. He was mentored by Robert, a Research Scientist from Wyoming Public Health Lab. After learning the basics of Nextflow and nf-core, they worked on a metatranscriptomics pipeline that simultaneously characterizes microbial composition and host gene expression RNA sequencing samples. As a part of this process, they used nf-core modules that were already available and developed and contributed new modules to the nf-core repository. Ícaro found having someone to help him learn and overcome issues as he was developing his pipeline was invaluable for his career.
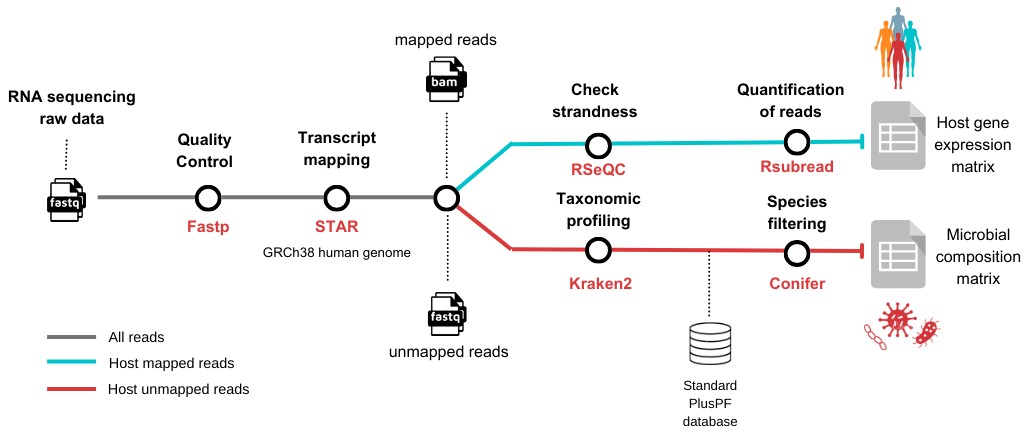
Metro map of the phiflow workflow.
Lila Maciel Rodríguez Pérez, from the National Agrarian University in Peru, was mentored by Priyanka Surana, a researcher from the Wellcome Sanger Institute in the UK. Lila and Priyanka focused on building and deploying Nextflow scripts for metagenomic assemblies. In particular, they were interested in the identification of Antibiotic-Resistant Genes (ARG), Metal-Resistant Genes (MRG), and Mobile Genetic Elements (MGE) in different environments, and in figuring out how these genes are correlated. Both Lila and Priyanka spoke highly of each other and how much they enjoyed being a part of the program.
Luisa is an MSc. student studying computational biology in the Computational Biology and Microbial Ecology group at Universidad de los Andes in Colombia. She was mentored by Gisela Gabernet, a researcher at Yale Medical School. At the start of the program, Luisa and Gisela focused on learning more about GitHub. They quickly moved on to developing an nf-core configuration file for Luisa’s local university cluster. Finally, they started developing a pipeline for the analysis of custom ONT metagenomic amplicons from coffee beans.
Natalia Coutoné is a Ph.D. Candidate at the University of Campinas in Brazil. Her mentor was Marcel Ribeiro-Dantas from Seqera. Natalia and Marcel worked on developing a pipeline to identify relevant QTL among two or more pool-seq samples. Learning the little things, such as how and where to get help was a valuable part of the learning process for Natalia. She also found it especially useful to consolidate a “Frankenstein” pipeline she had been using into a cohesive Nextflow pipeline that she could share with others.
Raquel Manzano is a bioinformatician and Ph.D. candidate at the University of Cambridge, Cancer Research UK Cambridge Institute. She was mentored by Maxime Garcia, a bioinformatics engineer at Seqera. During the program, they spent their time developing the nf-core/rnadnavar pipeline. Initially designed for cancer research, this pipeline identifies a consensus call set from RNA and DNA somatic variant calling tools. Both Raquel and Maxime found the program to be highly rewarding. Raquel’s presentation about the rnadnavar pipeline and her experience as a mentee from the 2023 Nextflow Summit in Barcelona is now online.
We are thrilled to report that the feedback from both mentors and mentees has been overwhelmingly positive. Every participant, whether mentor or mentee, found the experience extremely valuable and expressed gratitude for the chance to participate.
“I loved the experience and the opportunity to develop my autonomy in nextflow/nf-core. This community is totally amazing!” - Icaro Castro
“I think this was a great opportunity to learn about a tool that can make our day-to-day easier and reproducible. Who knows, maybe it can give you a better chance when applying for jobs.” - Alan Möbbs
Thanks to the fantastic support of the Chan Zuckerberg Initiative Diversity and Inclusion grant, Seqera, and our fantastic community, who made it possible to run all three rounds of the Nextflow and nf-core mentorship program.